Epigenetics takes us back to the Galápagos

Although not the most important among the many different animals studied by Charles Darwin during his amazing journeys on board the Beagle, the little finches from the Galápagos Islands have become one of the most popular representatives of Darwin’s theory of natural selection. They embody the process of speciation forced by environmental conditions, in the form of a bunch of different but very close species of birds that differ substantially depending on the particular island where they originated. With the pass of time, this group of animals has even been named as “Darwin’s finches”. Regardless of their role in the development of the theory of evolution (in the end they were just an example out of dozens), these species have continued being studied, and their genomes scrutinized at the molecular level in order to try to understand the particularities of the speciation process. At the present time, they have suddenly found themselves in the middle of a strange conflict involving opinions from both classical darwinists and revolution-eager epigenetists that claim for a rethinking of the whole evolutionary theory; to make things even funnier, our feathered friends have even been said to be a living proof of intelligent design. But let us first talk about the science, the facts, the data; and after that, we can discuss a little bit about those multiple and quite imaginative interpretations.
Epigenetics is the branch of modern genetics that studies a particular kind of changes in gene expression: those mediated by chemical modifications of DNA and its protein components, that do not affect the nucleotidic sequence. These changes are tightly interconnected to environmental influences. Thus, epigenetic mechanisms have since long been proposed as mediators of particular adaptive features of cells, and in extension, they might putatively influence the evolution of a whole organism in response to outer conditioners. To date, no definitive data have corroborated or dismissed this possible role of epigenetics, and this is why Skinner and collaborators decided to study Darwin’s finches from a double genetic perspective, comparing the diversification of the different finch species with the specific patterns of genetic and epigenetic changes.
The result of this work has just been published in Genome Biology and Evolution, under the slightly sensationalist title “Epigenetics and the evolution of Darwin’s finches” 1. Skinner and his team chose to compare the pattern of copy number variations (CNVs), which are repetitions of genetic sequences that differ between closely related species in their number, and have been related to phenotypic variation and speciation processes; then, they chose a particular kind of epigenetic modification, DNA methylation, a chemical alteration of concrete DNA components that affects gene expression, inhibiting the activation of concrete genes and even promoting the alteration of the global structure of the genome. In short, methylation and demethylation constitutes a pattern of ON and OFF signals for gene activation that interestingly can be transmitted to the offspring, and therefore, it harbors the potential to act as an important factor in evolution. The question to ask was: which kind of modifications in the finches’ genomes, the “only genetic” or the “only epigenetic”, showed a pattern of divergence that could correlate with the phenotypic divergence observed in the diversity of Darwin’s finches? It might seem difficult to compare things in which evolution during millions of years has to be tested; in these cases, researchers have to choose a “standard” DNA sequence, to be used as model to compare with the DNA sequence of the rest of species. When comparing DNA or protein sequences, the study of their similarities and differences at the molecular level reflects the millions of years of evolution from their common ancestors (we saw a similar example in the study of eukaryotic kinetochores).
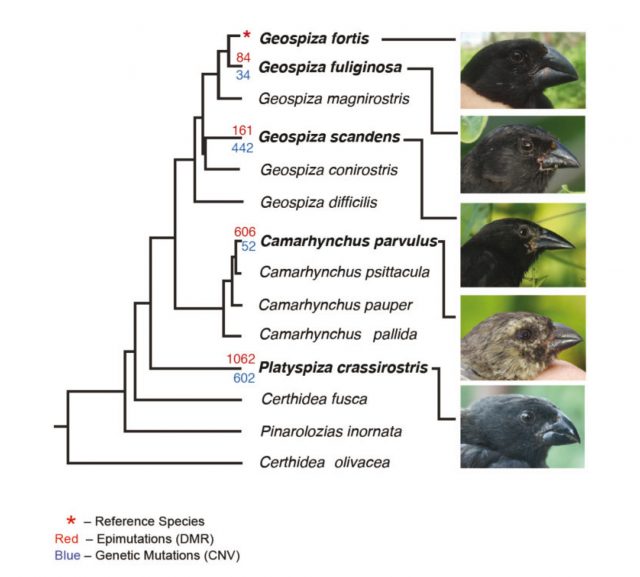
The chosen standard was the genome of Geospiza fortis, the reference species. The experiments consisted in purifying genomic DNA from blood samples and comparing one to another using the technology of hybridization in DNA arrays, in parallel to techniques of chromatin immunoprecipitation to assess the methylation patterns. Simplifying, CNVs were mapped and compared between species, and the same was done with DNA methylation patches. Who won this competition? Well, the results were clear: the genomic repeats were much more stable than the methylation pattern: in other words, as finches species diverged from a common ancestor, the methylation patterns also got more and more different, whereas the CNVs remained more or less the same. This particular variations of epigenetic marks paralleled the branching of finch species into phenotypically different adaptations (the shape of the beak has always been the most famous and obvious feature to pay attention to). To gain insight into the possible functional relevance of this finding, the researchers analyzed the presence of this epigenetic variations in particular molecular pathways; interestingly enough, the regions with the highest frequency of variation for methylation patches were always located in the vicinities of genes associated with avian evolution (for instance, some of those genes have been related with the bone morphogenetic process of the beak development).
The data offer a clear scenario: methylation processes are specific of each species of Darwin’s finches, whereas the variations in length of repeated regions are more conserved. Does this mean that epigenetic variations drove the speciation process in this group of birds? Probably, this is a pretty bold assessment. First, because only a pair of genetic features have been studied; second, because even though epigenetic patterns are there today, into the finches’ genomes, it is truly difficult to ascertain if they have been there for generations, affecting gene expression and provoking the fast changes that made those species different to one another. They could just be the manifestation of environmental differences that modify the genomes, with no significant effect on the evolutionary process.
It could be better argued that there is really no true sense in opposing “genetic” versus “epigenetic”. Epigenetic modifications are, in the end, no more than another kind of genetic variance; they sum up to the rest of origins of variation for gene expression, and must be fixed by natural selection, during generations, always under the influence of the environment. In the last years, many details of epigenetic mechanisms are being unveiled, that uncover a vast spectrum of possibilities and fine regulations of gene expression, in many occasions tightly linked to changes in the surrounding of the cells. Following these evidences, some scientists are claiming that the role of epigenetic modifications in evolution is more important that initially thought 2. But the notion is not as revolutionary as it may seem; genetic changes have always been known to be affected by the environment. Gene expression is subjected to changes in the exterior, and the way a cell responds depends always in its genetic programing, which by the way, includes its epigenetic patterns (the proteins that modify epigenetic marks are also coded into the genome). The data provided by this work provide valuable evidences of the participation of epigenetic regulation in configuring the phenotypic aspect of organisms during their adaption, and probably reflect an especially relevant role in radiation processes like the one observed in Darwin’s finches. However, their capacity to drive evolution has yet to be proved. Specially challenging will be the demonstration that these modifications are really influenced by external stimuli, and more important, that they are fixed into the genome and passed from generation to generation long enough as to culminate in the configuration of a different species. An interesting test would be to breed one of these finch species in an environment different to its original island, and then check if the methylation pattern varies, and how. But this is usually the problem with evolutionary studies: testing some hypothesis is quite difficult, unless you dispose of generations of researchers eager to wait and see the result of their ancestor’s preliminary studies.
Finally, a curious fact: among all the polemics and controversies usually found around epigenetics and the fashionable topic of “nature” versus “nurture” debates, there is one that impersonates the most imaginative and unexpected. The very same paper we have just reviewed today is being raised by the defenders of intelligent design as the definitive proof that the actual theory of evolution is wrong. They see in epigenetic mechanisms a way to explain the Earth’s diversity that permits the assumption that all animals on our planet were created independently, not deriving from previous ancestors after millions of years of change accumulation. In all, this is a nice example of the differences between science and faith; taking a series of facts and adapting them to a previously assumed scenario, instead of collecting information and asking opposite questions in order to find which answer supports the observed data.
Probably Charles Darwin, a humble naturalist as he was, would be surprised to know that with all our modern knowledge of biology, we still follow the clues he first pointed at by observing these little animals and their habitats. He never gave the finches from the Galápagos as much attention, and he surely would be dazzled to find that they are being thoroughly examined until the deepest cell and, surprisingly, claimed by creationists as the definitive nail in the coffin of Darwinism.
References
- Skinner M.K., M. M. Haque, E. E. Nilsson, J. A. H. Koop, S. A. Knutie & D. H. Clayton (2014). Epigenetics and the Evolution of Darwin’s Finches, Genome Biology and Evolution, 6 (8) 1972-1989. DOI: http://dx.doi.org/10.1093/gbe/evu158 ↩
- Laland K, Uller T, Feldman M, Sterelny K, Müller GB, Moczek A, Jablonka E, Odling-Smee J, Wray GA, Hoekstra HE, Futuyma DJ, Lenski RE, Mackay TF, Schluter D, Strassmann JE. (2014) Does evolutionary theory need a rethink? Nature 514(7521):161-4. doi: 10.1038/514161a. ↩
2 comments
[…] Darwinen txontei buruz guztia esanda zegoela zirudienean, darwinista klasiko eta epigenetiko iraultzaileen arteko eztabaida dugu pil-pilean. Eta tartean, diseinu adimentsuaren aldekoak muturra sartzen. Carlos Romá-Mateoren artikuluan ditugu guztiak: Epigenetics takes us back to the Galápagos. […]
[…] Cuando parecía que ya se había dicho todo sobre los pinzones de Darwin, resulta que ahora se encuentran en medio de un debate entre darwinistas clásicos y epigenetistas revolucionarios; con los partidarios del diseño inteligente metiendo baza. Carlos Romá-Mateo […]