Living beings: systems all the way back to their chemical origins
Living beings: systems all the way back to their chemical origins
Author: Kepa Ruiz-Mirazo, Permanent Researcher, University of the Basque Country (UPV-EHU)
The effort of mapping ignorance in biology is especially frustrating –or, if the mood and attitude of the scientist afford it, especially motivating– because solid, definite answers remain minimal compared to the amount of open issues and uncertainties that new discoveries in this field bring about. There is no doubt that our knowledge about the living world increases every day, but evidence suggesting gaps in our understanding, the possibility that even the simplest cells on Earth (bacteria, like Mycoplasma) might be too complex for our limited cognitive capacities, accumulates much more rapidly. In such unfair epistemological conditions, in which the sphere of the unknown seems to grow always larger and faster than the known, a common strategy among biologists has been to give in, somehow, and concentrate on the study of specific molecular mechanisms underlying particular aspects of the behavior of certain organisms.
This analytic-descriptive strategy has been immensely fruitful, though. The corpus of knowledge produced in that way over the years is not only amazing in terms of size and level of detail, but also of fundamental importance for our general comprehension of the molecular bases on which all biological phenomena stand. Quite surely, there was no other way: it was necessary to start from the particulars in order to assess, later on, whether generalizations or common principles were also feasible within the living domain. Furthermore, without that precise molecular knowledge biology would lack most of its current transformative power: biotechnology simply could not have developed. So one of the central tasks of biological sciences is to continue accumulating data about the huge diversity of living beings in the biosphere and invent new techniques to extract more and more accurate information on how they operate molecularly. However, this will lead us nowhere unless we find suitable guidelines to interpret and integrate all that data. Luckily, a significant part of the scientific community has realized this now, and the emergence and progressive consolidation of disciplines like ‘systems biology’, ‘bioinformatics’ or ‘synthetic biology’ clearly reflects the present urge for having better criteria and insights about how biomolecules interact within complex modules or networks; about how they get functionally coupled in the constitution of complete living beings.
Nevertheless, this may be insufficient to decipher what we are missing — at any rate, to be able to draw a minimally reliable map of our ignorance. Despite all the novel and highly promising scientific enterprises of 21st-century biology devoted to integrative-synthetic tasks, living systems will probably remain a mystery until we make real progress in explaining how they came about. In fact, the problem of origins is intrinsically connected with the one of determining what life consists in, like Oparin already remarked in his days. On these lines, even if the question about the inert-living transition is still far from being solved, chemistry has recently started setting up the stage for a proper approach to it. Up to now, the field of ‘prebiotic chemistry’ was mainly concerned with discovering abiotic reaction pathways that could lead to the key organic molecules present in every biological organism on this planet: RNA, DNA, proteins, lipids, sugars… Very few attempts had been made, though, to put these biomolecules –or, more reasonably, their corresponding precursors– together. It was simpler, in a first approximation, to go step by step and deal with each type of molecule at a time. As a result, the research field divided into different schools, each of them embracing a hypothesis of work that put a specific type of biomolecule as the primary one: the ‘RNA world’, the ‘lipid world’, the ‘protein world’… And the emphasis was therefore placed on determining the most plausible chemical reaction routes that could account for the formation of each of those families of biomolecules, as they are found in extant living beings — but considered separately.
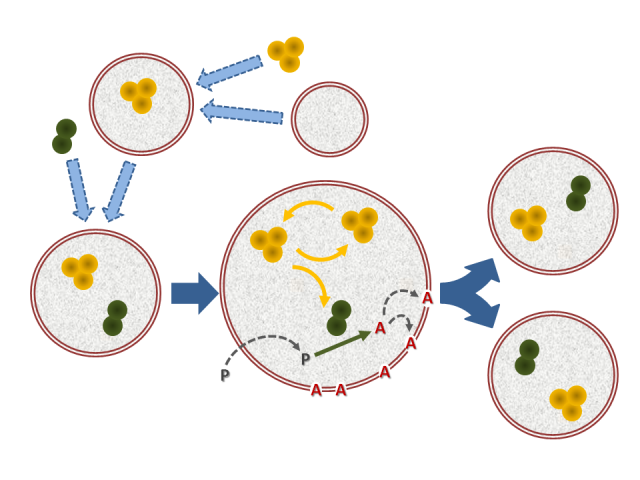
However, this has little to do with biological reality. Cells, as everybody knows, are made of those complex organic molecules; yet the latter come to existence and play a role by virtue of an intricate web of dynamic inter-relationships and transformation processes, which in practice holds all of them together. Remarkably, most biomolecules (except for DNA) have relatively short ‘lifetimes’, much shorter than the lifetime of the collective entity they integrate. Each living organism is a molecular factory in continuous turnover, whose far from equilibrium activity is maintained precisely thanks to the generation of all those structures that are required to build the factory up and keep it running. Without the factory the various pieces, their sheer existence and purpose, just would not make sense: each is produced and has a function in the context of the activity of all the others. If this fact is seriously assimilated, it becomes obvious that the first appearance of something resembling a living being could not be thanks to molecules of a single type. It would be easier to study if it were in that way, no doubt, but life is hard — also for researchers in the field of origins.
Fortunately (for scientists in general, but for biologists in particular), chemistry is moving forward, taking very important steps to face the challenge of complexity upfront. The newly created field of ‘systems chemistry’ is developing at present the techniques and skills to study in detail heterogeneous mixtures of molecules: big libraries of chemical components in interaction; processes of synchronization among different reaction dynamics; emergence of spatial and temporal patterns; the spontaneous generation of macroscopic correlations among millions of microscopically reacting elements of diverse species, in diverse media; combinations of self-assembling quasi-equilibrium supramolecular structures with far-from-equilibrium self-organizing processes. In brief, what was just a dream for chemists in a very recent past is becoming a real possibility today, also because they have the suitable technology in their hands now: micro- and nano-fluidic devices, dynamic combinatorial libraries (DCLs), high-throughput sequencing and microarrays, powerful parallel computing for theoretical simulations,…
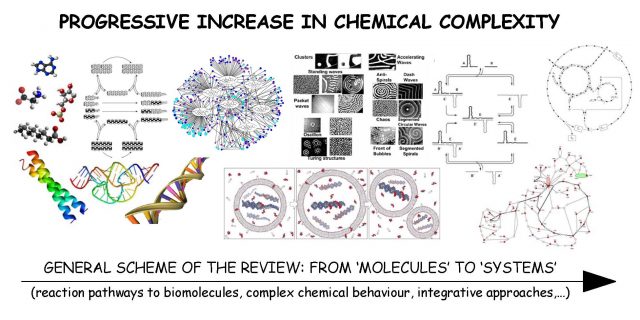
All this will have long-lasting consequences and enormous applicability in many areas, but one of the most direct beneficiaries will be the field of ‘origins of life’, as we recently highlighted in an extensive review 1. Scientists interested in the chemistry-biology gap will find, at last, a natural bridge between the two: truly systemic approaches linking the complexity of the first with the hyper-complexity of the second. Research shortcuts or excursions based on the properties of single types of molecules may be very important in various respects, practical and theoretical, but are not helpful if the aim is to cross that river, deep and wide, that still separates us from our chemical roots. In a similar way as the building of a real, long and firm bridge requires various materials and mechanisms of integration, our metaphorical bridge requires heterogeneity in composition and multiple dynamic and functional couplings. Nature became a “perfect queen” in these affairs long ago: the integration of diverse molecular components performing different tasks within a complex organization surely started at a proto-cellular stage, long before natural selection pushed the birth of more robust, full-fledged living cells — made of DNA, RNA and proteins. Science is just getting ready these days to be apt to formulate some relevant questions about the process of biogenesis within this alternative, still underexplored scenario; and hopefully also answer part of them, so that the journey to map our ignorance continues.
More info: COST Action on systems chemistry
References
- Ruiz-Mirazo, K., Briones, C. & de la Escosura, A. (2014): Prebiotic Systems Chemistry: New Perspectives for the Origins of Life. Chemical Reviews 114: 285-366. DOI: 10.1021/cr2004844 ↩
3 comments
[…] mappingignorance.org […]
[…] sus componentes (con la excepción de su genoma) dentro del equilibrio cinético en el que viven. La biología y la química de sistemas investigan sobre ello pero, por el momento, aunque dispusiéramos de una cantidad ilimitada de […]
[…] los planteamientos y metodologías característicos de una disciplina emergente conocida como química de sistemas, de la que son pioneros investigadores como Günter von Kiedrowski o Sijbren […]