Molecular Detectives: discovering new ion channels (II)

The identification of previously unknown proteins is a difficult task and often requires to follow unconventional thinking. In my previous post, I have described how the TRPV1 ion channel (formerly known as the capsaicin receptor) was discovered by combining the construction of a cDNA library from primary afferent neurons (DRG neurons) and the functional selection of clones using fluorescent dyes as a main tool. In that case however, the vanilloid capsaicin turned out to be exquisitely selective for TRPV1, thus enormously facilitating the process of choosing the right clones. How would we identify an ion channel whose gating relies upon a rather vague stimulus such as physical forces?
Our body needs to detect mechanical forces such as vibration, sound waves or stretch to feel touch, hear sounds and even for the unconscious sensation of blood flow. For instance, auditory receptor cells contain a structure called sterocilia interconnected by tip links. When sound waves mechanically bend the esterocilia, the tip links are pulled leading to the activation of mechanically-sensitive ion channel expressed within them.
Is therefore important to find out genes encoding mechanically-activated (MA) ion channels not only in order to better understand how those physiological processes work but also to be able to link them to a potential causative pathological condition. Unfortunately, this has been proven an arduous task. In spite of the first electrophysiological evidence suggesting the existence of MA ion channels dating from 1974, there is controversy as to whether the proposed ion channels directly confer mechanosensitivity, so far. A comprehensive review on that matter by Sanjeev S. Ranade et al is available for further reading 1 .
Yet a paper published in 2010 in the prestigious journal Science by the group from Ardem Patapoutian successfully identified two previously unknown “components of MA cation channels” 2. They called them “components” because at such early stage, they had no proof of them being actual pore-forming membrane receptors (ion channels). How could they accomplish such task? As their interest was mainly focused in mechanical pain detection (similarly to the group who discovered TRPV1 ion channel), they first aimed to find a cell line showing a robust mechanically-mediated electrical response comparable to that obtained from DRG neurons. The reason why not to use primary afferent neurons as starting material may rely on a number of advantages, including genetic homogeneity across cells, growth speed and avoid sacrificing laboratory animals throughout the screening process, which can end up being quite lengthy.
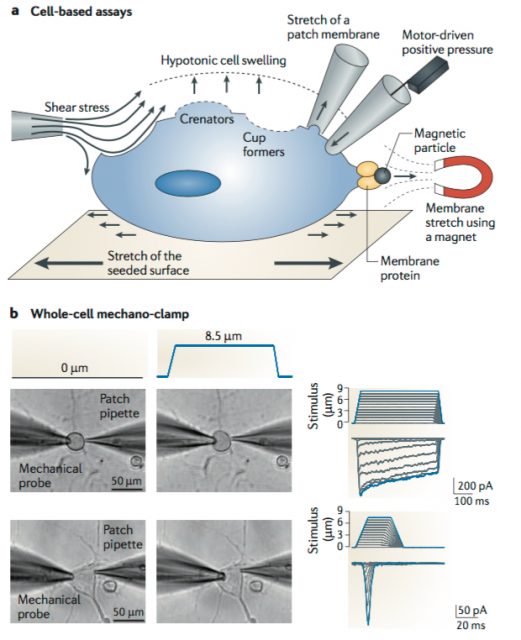
After functional assessment of several cell lines with patch-clamp, they found applying mechanical pressure onto a mice neuroblastoma-derived cell line (N2A line) led to the activation of an electrical current with the expected biophysical properties. To do so, they utilized a piezo-electrically driven glass probe which allowed them to precisely control the amount of pressure provided to the cells (Figure 1). Next, they used an Affymetrix microarray (essentially, a solid surface including a collection of DNA probes) to look for transcripts that were enriched in N2A and create a list of potential MA candidates based on the following criteria. The protein of interest should have a predicted structure comprising at least two transmembrane domains (minimum requirement for a membrane protein to be considered an ion channel) and from those, they would choose either cation channels (based on the known properties of MA currents) or proteins with unknown function. They ended up with 73 potential targets meeting the criteria.
The subsequent smart move was to knockdown each of the 73 gene candidates by synthesizing small interfering RNA (siRNA) complementary to them. This technique is used to prevent the translation of proteins by binding of the siRNA to the complementary messenger RNA at specific sequences, resulting on its degradation. They would just have to inject each of these siRNAs into the N2A cell line and apply mechanical pressure, looking for the one blocking the appearance of the MA current. After screening all 73 genes (unfortunately, the last one turned out to be the hit), they found the Family with sequence similarity 38, member A (Fam38A) gene encoded a previously uncharacterized protein.
The newly found ion channel was named Piezo1 and is evolutionarily conserved among animals and plants, although their sequence is not homologous to any other ion channel family, suggesting it was the founding member of a new class. In addition, they identified another member in vertebrates (Piezo2) which was found to be expressed in a smaller proportion of DRG neurons (20%). Both members seem to be non-selective cationic channels and were activated upon mechanical stimulus, but they do show slightly different biophysical properties. Computational models and biochemical studies predict an unusual topology with 18 to 38 transmembrane segments, meaning they may represent the largest ion channel complex identified, so far. Most notable, expression of the Fam38A gene into naive cells confers mechanical sensitivity, indicating that Piezo1 is mechanically gated itself, without the need of other accessory elements.
Since their discovery, many laboratories have focused in deciphering the physiological roles of Piezo. Piezo1 play a key role in the development of the mouse vascular system and when knocked down specifically in red blood cells from adult mouse, leads to increased fragility and dehydration. Meanwhile, Piezo2 has been established as the primary detector of light touch in mammals and mutations have been linked to several diseases, including the Gordon syndrome, Marden-Walker syndrome and Arthrogryposis. These disorders often lead to stiffness and impaired mobility of joints from distal extremities (knees, elbows, wrists and ankles).
In summary, the identification of new proteins not only serves the purpose of giving credit to the scientists involved, but also to make great strides in basic research. It is therefore important not forgetting that even though promoting applied research does lead to great progress, basic research unlocks new unanticipated and exciting paths.
References
- Ranade SS, Syeda R, Patapoutian A. Mechanically Activated Ion Channels. Neuron. 2015 Sep 23;87(6):1162-79. doi: 10.1016/j.neuron.2015.08.032. Review. Erratum in: Neuron. 2015 Oct 21;88(2):433. ↩
- Coste B, Mathur J, Schmidt M, Earley TJ, Ranade S, Petrus MJ, Dubin AE, Patapoutian A. Piezo1 and Piezo2 are essential components of distinct mechanically activated cation channels. Science. 2010 Oct 1;330(6000):55-60. doi: 10.1126/science.1193270. ↩
3 comments
Excellent (!) article, thanks a lot for sharing.
Dear Mathieu,
I’m glad you liked it. This is a fascinating field that has been growing a lot in the last years. Will try to update when new progress is achieved.
[…] El proceso para descubrir nuevos canales iónicos mecánicos es digno de Sherlock Holmes. Sergio Laínez en Molecular Detectives: discovering new ion channels (II) […]