The rise of the superbugs
The rise of the superbugs
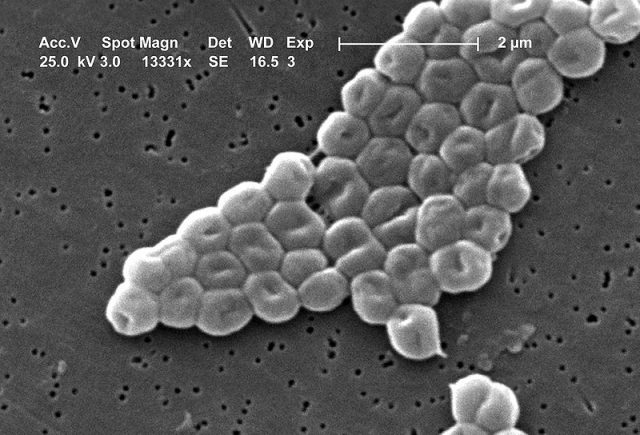
Our society will be facing a number of health-related challenges in the near future, partly as a consequence of our own practices. Perhaps the best known examples are the (severe) health issues linked to smoking (not just to smokers, but also their children via epigenetic changes) or the abnormally high intake of carbohydrates, with the latter one becoming an ever-growing burden for the public health system (and entirely preventable). Regrettably, these are not isolated issues and there is yet another fast-growing -and potentially worse- problem, which is the rise of antimicrobial resistance (AMR) pathogens, the so-called superbugs.
Antibiotic prescription has revolutionized medicine since it was serendipitously discovered by Fleming back in 1928, but it has come at a price. Despite the tendency to unnecessarily prescribe antibiotics by physicians –with ~30% of antibiotics prescribed in US being dispensable – and the debate as to whether this has led to a faster acquisition rate of resistance by bacterial strains, the genetic versatility of these unicellular organisms allows them to rapidly develop resistance to many antibiotics currently in use. Still, the acquisition of resistance varies depending on the bacterial species and the antibiotic under consideration. As an illustrative example, Spain had in 2015 a 25-50% range of Escherichia coli invasive isolates with resistance to fluoroquinolones, while the percentage of the same bacteria resistant to third-generation cephalosporins was 10-25%, suggesting we are not in a national health emergency, but some sort of strategy should be in place. Perhaps the most pressing issue is the occurrence of nosocomial infections, which are those acquired by different means (e.g. contaminated equipment, staff gowns, among others) in hospitals or related health care facilities, accounting for at least 2.5 million new healthcare-associated infections per year just in Europe.
Mutations in bacterial own genes or horizontal acquisition of exogenous mobile genetic elements between bacteria represent the main mechanisms explaining the acquisition of resistance. At the functional level, resistance involves enzymatic degradation of antibiotics, modification of antibiotic protein targets or changes in membrane permeability to them. One of the “usual suspects” involved in nosocomial infections is the gram-negative Acinetobacter baumannii, which causes thousands of deaths every year –via pneumonia or bloodstream infections- and has been tagged as “serious threat” by the US Centers for Disease Control. The underlying reason seems to be an abnormally fast rate of resistance acquisition through horizontal gene transfer (HGT), not being clear how they can achieve such high transfer rate.
In order to shed light on that matter, Cooper et al decided to study the well-known relative Acinetobacter baylyi, which is arguably not the best model to use for comparison purposes -as is a soil bacterium lacking clinical relevance- but as a basic research approximation is quite interesting. They co-cultured A. baylyi and E. coli containing green fluorescence and AMR genes against kanamycin on a microfluidics device allowing single cell monolayer visualization 1. As both bacteria began to grow, they observed lysis of E. coli accompanied by the appearance of green fluorescing A. baylyi, indicative of HGT (Figure 1). Likewise, when Kanamycin was added to the culture, these brand new green Acinetobacter kept growing, confirming the AMR genes had been transferred too (Figure 1). Interestingly, HGT was dramatically reduced when the ability of A. baylyi to kill was inhibited, indicating the high HGT rate relies upon a cell killing-dependent mechanism. However, when killing-dependent HGT happens, the newly acquired genes can be uptaken by nearby non-killing cells. That means that the presence of different microbes together further facilitates HGT, and that is not good news, as different bacterial strains cohabitate in hospitals.
Finally, they narrow-down conditions at which maximal HGT rates happen. These include: The need for direct contact between predator and prey cells, and high predator to low prey density (especially at the beginning of the experiments). These observations represent valuable information that should enable scientists to find more efficient ways to fight AMR, and provide guidance to assess if similar transfer rates and mechanism are common to other bacterial strains tagged as “serious threats” too.
Furthermore, authorities should envisage other types of measures which should then be efficiently implemented and strictly followed by healthcare professionals. Some of them may include: The prevention of infections and spread of resistant bacteria (hygienic measures, especially in hospital environment); continuous tracking of resistant bacteria (development of commercial kits allowing the identification of hundreds of bacterial strains); optimize the use of antibiotics by reducing their prescription and foster the development of new types of antibiotics and fast diagnostic tests.
References
- Cooper RM, Tsimring L, Hasty J. Inter-species population dynamics enhance microbial horizontal gene transfer and spread of antibiotic resistance. Elife. 2017 Nov 1;6. pii: e25950. doi: 10.7554/eLife.25950. ↩
1 comment
[…] eta obesistatea baino gehiago ere. Zelan garatzen da erresistentzia hori, baina? Sergio Laínez The rise of the superbugs […]