The ribosome world hypothesis
The ribosome world hypothesis
The ribosome may be a missing link in the evolution of life. This suggestive proposal has been published by Meredith Root-Bernstein, Oxford University, UK, and her father Robert Root-Bernstein, Michigan State University, USA, in the Journal of Theoretical Biology 1. Their hypothesis is that primordial ribosomes were self-replicating intermediates between the prebiotic world and the first cellular form of life, LUCA (Last Universal Common Ancestor).

Life requires three polymers: DNA, RNA, and proteins. However, the current thinking on the emergence of life considers DNA as a secondary product. The two dominant hypothesis for the origin of life are the RNA world (genetics-first) 2 and the compositional replication (metabolism-first) models 3. Precursor organic molecules (sugars, nucleotides, amino acids, etc.) were readily available in the primordial environment through inorganic, prebiotic chemical reactions. Hence, the co-evolution of nucleic acids and polypeptides is almost certain. Therefore, pre-cell entities only made of RNA and proteins are the most promising candidates to pre-date LUCA.
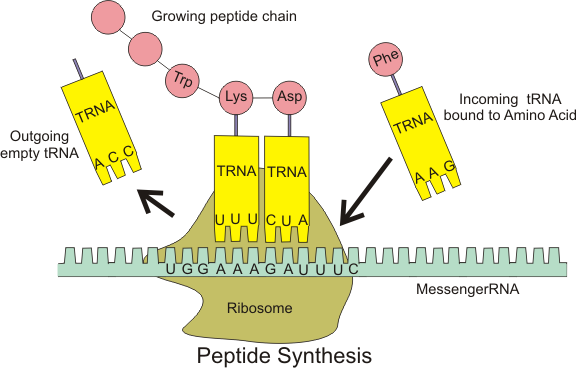
Ribosomes are made of ribosomal RNA (rRNA) molecules and proteins. They are key cell organelles since they are responsible for protein synthesis (translation). Ribosomes link amino acids together in the order specified by messenger RNA (mRNA) molecules. Amino acids are carried to the ribosome by transfer RNA (tRNA) molecules matching the genetic code, which uses three-nucleotide sequences to encode every amino acid. A protein, the enzyme RNA polymerase, is responsible for the transcription of particular DNA segments (genes) into mRNA molecules.
The ribosome world hypothesis of the Root-Bernsteins has similarities with the endosymbiotic theory, or symbiogenesis, advocated by Lynn Margulis. In such a theory the key organelles of eukaryotes originated as symbiosis between separate prokaryotes. For mitochondria, chloroplasts and other plastids there is some molecular and biochemical evidence suggesting the hypothesis. In the ribosome world, in addition to ribosomes, there must be other pre-organelles, like acidocalcisomes (the only cellular organelles conserved between prokaryotic and eukaryotic organisms which are implied in osmoregulation). However, the new paper only focuses on the ribosome genome.
The pre-ribosomes must be self-replicating. However, ribosomes do not carry genetic information, being their rRNA molecules only structural components. Reference [1] studies the possibility that rRNA encoded the tRNAs and proteins necessary to ribosomal function. In such a case, pre-ribosomes may have the capability of genetically encoding their own transcription and translation machinery. To explore the possibility, modern bioinformatics algorithms for sequence alignment (like LALIGN and BLAST) could be applied. For concreteness, the new paper considers the three rRNA sequences of the bacterium Escherichia coli (5S, 16S, and 23S).
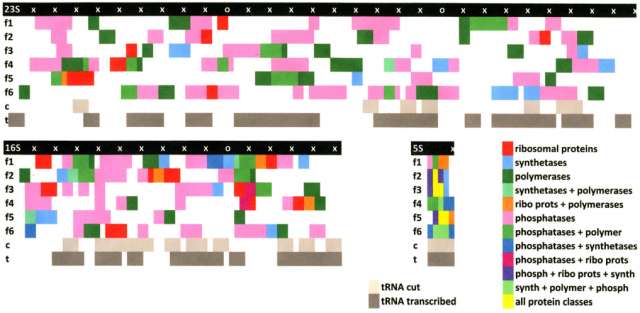
The results of the Root-Bernsteins, in their opinion, clearly favor their hypothesis, that the ribosome may have been a primordial self-replicating entity, over the alternative hypothesis that rRNA contains no genetic information or that it contains random genetic information. All genetically encoded tRNAs for all twenty amino acids are encoded (with overlapping) in the 16S and 23S rRNAs. In fact, it would be possible to generate all the tRNA sequences by cutting or editing the rRNA itself into appropriate fragments. These overlap encoding is common in mitochondria and animals (metazoans), being functional at least in mitochondria. But they are absent in unicellular organisms like E. coli or yeast. However, yeast apparently retains the vestigial mechanisms for tRNA editing to process these overlaps functionally 4.
The possibility that E. coli rRNA also encodes proteins is more difficult to study since the protein similarities found may have occurred by chance. The analysis of the Root-Bernsteins show some promising evidence for their hypothesis, but it is not conclusive. For example, rRNA encodes possible polyribonucleotide polymerase fragments that could potentially have participated in the replication of RNA. They also encode some fragments of proteins essential for energy storage and transduction. But in my opinion more evidence is required to claim that these results are evidence that primitive ribosome-like entities may have encoded the basic elements of the energy metabolism required to drive ribosomal function. Moreover, further experiments are required to verify whether the ribosome-related protein sequences rRNA-encoded in E. coli are truly functional.
Bacterial ribosome translating RNA into protein. Source: Venki Ramakrishnan’s group at the LMB in Cambridge, UK.
The new paper provides some hints about how and why DNA storage of genetic information may have evolved. The Root-Bernsteins guess that the synthesis of DNA and the specialization of RNA into ribosomal, messenger, and transfer are natural by-products of RNA replication. Chemical evolution by natural selection prefers DNA to RNA since it is a more stable template for ribosomal survival. They suggest that “selfish ribosomes” may be the origin of genes, paraphrasing “selfish gene” introduced by Richard Dawkins.
In summary, the ribosome world proposed by the Root-Bernsteins is supported only by the results of RNA and DNA sequence alignments in E. coli. But several testable predictions follow, for example, that other bacteria must have rRNA–tRNA regions and protein modules encoded in rRNA very similar to those shared by the E. coli, or that some of the tRNAs encoded in rRNAs retain functionality. Further research is required to contrast these predictions.
References
- Root-Bernstein M. (2015). The ribosome as a missing link in the evolution of life, Journal of Theoretical Biology, 367 130-158. DOI: http://dx.doi.org/10.1016/j.jtbi.2014.11.025 ↩
- Marc Neveu, Hyo-Joong Kim, Steven A. Benner, “The ‘Strong’ RNA World Hypothesis: Fifty Years Old,” Astrobiology 13: 391–403, Apr 2013; doi:10.1089/ast.2012.0868 ↩
- Axel Hunding et al., “Compositional complementarity and prebiotic ecology in the origin of life,” BioEssays 28: 399–412, Apr 2006; doi:10.1002/bies.20389 ↩
- Jens Schuster, Heike Betat, Mario Mörl, “Is yeast on its way to evolving tRNA editing?,” EMBO Reports 6: 367–372, Apr 2005; doi:10.1038/sj.embor.7400381 ↩
3 comments
[…] (aunque en inglés) Francisco R. Villatoro, “The ribosome world hypothesis,” Mapping Ignorance, 27 Feb 2015; el artículo es M. Root-Bernstein, R. Root-Bernstein, “The ribosome as a missing link […]
[…] (aunque en inglés) Francisco R. Villatoro, “The ribosome world hypothesis,” Mapping Ignorance, 27 Feb 2015; el artículo es M. Root-Bernstein, R. Root-Bernstein, “The ribosome as a missing link […]
[…] Francisco R. Villatoro, “The ribosome world hypothesis”, Mapping Ignorance, 27 Feb 2015. […]