The flick of a switch controls the fate of human parasites
The flick of a switch controls the fate of human parasites
The advances in molecular biology and the so called post-genomic era, have improved significantly the fight against many human diseases, in some cases almost leading to their eradication. However, there are still regions of our planet were people suffer from infections and other causes of mortality which are easily avoided in the most developed countries. Research on these pathologies is therefore critically important. In this article we will focus on two parasites which are responsible for two human diseases, malaria and toxoplasmosis.
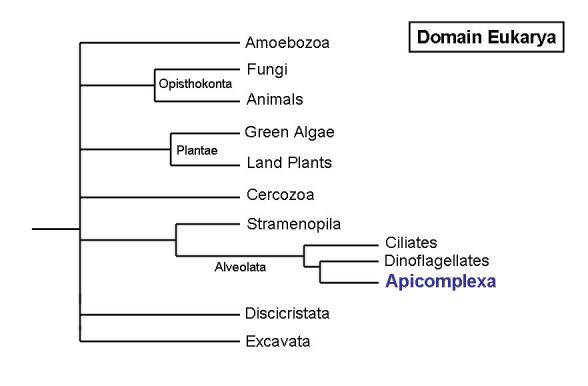
Malaria infections are terribly rife in many African countries; toxoplasmosis, although generally non-pathological for the majority of the population, becomes a great risk for immunocompromised patients and it is an important cause of abortion and newborn malformations. Both diseases are caused by different infectious agents, both of which belong to an evolutionarily related group: the unicellular organisms known as apicomplexa. Apicomplexa are a group of unicellular, obligate parasites which form part of the higher group alveolata (Figure 1). Though evolutionarily related, the organisms causing malaria (species belonging to genus Plasmodium) or toxoplasmosis (genus Toxoplasma) show strikingly different life cycles and reproductive strategies.
Plasmodium parasites’ life cycle involves the development of different forms as it passes from a carrier organism (called the animal vector, a mosquito in this case) to the vertebrate where it fully develops. The organism suffers a total transformation in its different stages, from latent forms called oocysts, to the infective forms called schizonts that invade the mammalian red cells (Figure 2). Toxoplasma parasites are also present in latent forms called bradyzoites or proliferative, infective forms called tachyzoites. However, the latter develop inside vertebrate organisms. A curious course of the parasite evolution has ended up with cats being the definitive organisms where the reproductive life cycle of the parasite is fulfilled.
These huge differences in the ways of life of the parasites appear drastically reduced when we get to the molecular level. All organisms control the appearance, function and fate of their cells by means of a translational program which converts the instructions coded in the genome (the content of DNA, which remains immutable for all the cells of an organism) in a particular behavior of the cells. In these parasites, which are constituted by a single cell, the translational control of their genome results in a radical transformation of the whole organisms that facilitates the accomplishment of the difficult task implied in surviving in surroundings as different as the interior of a mosquito, or the muscular structures of a vertebrate. Along the way, the parasites must survive in hostile environments, reproduce, and at the same time fight the host’s immune system.
A recent review 1 by Min Zhang and collaborators, from The New York University School of Medicine and Indiana University School of Medicine and published in Eukaryotic Cell, presents a summary of the translational strategies and proteomic changes reported for Plasmodium and Toxoplasma parasites during their different life stages. Putting together a nice amount of data, the landscape that comes out from this review is that the most critical changes in the protein composition of the different life forms of both parasites focus in the translational control. We should specify what “translational control” refers to.
Translation begins when a set of proteins assembles with the ribosomes, macromolecular complexes that bind mRNA (messenger RNA, produced after transcription of DNA information into a more summarized set of instructions). One of the critical proteins in this translational process is the proteinacious factor named eIF2 (eukaryotic Initiation Factor), which binds ribosomes and other molecules in order to mediate initiation of translation (this group of proteins bound to eIF2 is then called “initiation complex”). But the construction of proteins in the cells is far more complex, and eIF2 activity is dependent of many factors: what signals from outside the cell are being received, which other proteins bind to eIF2 in a precise moment, and also what modifications affect eIF2 itself.
One of the most critical modifications of eIF2 is phosphorylation. This is a very extended mechanism by which proteins incorporate a molecule of phosphate into their structure, mediated by other regulator proteins known as kinases. This modification alters their behavior and may produce inactivation, among other changes. It is a reversible action counteracted by another type of proteins, phosphatases, which remove phosphate groups. In this case, eIF2 phosphorylation in humans produces a change in the preference for the mRNAs which are a target for eIF2 after the formation of the initiation complex.
Briefly, when eIF2 is phosphorylated, it activates a cascade of events that preferentially result in the construction of a set of proteins and molecules with the role of fighting cellular stresses. When this stress-response is prioritary, a large set of instructions (this means a set of untranslated mRNAs) for the normal cell behavior accumulates in the so-called stress granules, waiting for better conditions to start their conversion into a new profile of protein expression (expression profiles are the set of proteins which are created in a given moment in a concrete cell).

In the aforementioned parasites, there is a protein complex equivalent to the human eIF2. Interestingly, the authors of the review find a large set of published data that correlate the phosphorylation state of Plasmodium and Toxoplasma versions of eIF2 with critical changes in protein expression that result in the conversion of latent forms of the parasite into infective or reproductive ones. These changes are always triggered by the surroundings of the organisms, which articulate different stress conditions. For the parasite, the stress generated by a host immune response is what starts the chain of events that convert the initial infective form into a latent and resistant form, awaiting for a better moment or place to continue the life cycle. This complex metamorphosis is achieved by a change in protein expression mainly motivated by alterations in the phosphorylation of the translational master switch that becomes eIF2. For instance, the activity of one of the kinases regulating phosphorylation of eIF2 in Plasmodium, eIFK2, is up-regulated in the sporozoytes carried by the salivary glands of the mosquito: thus, eIF2 in its phosphorylated form does not accomplish translation of any mRNA related to the development of the parasite forms that infect the vertebrate liver. Once in the host organism, phosphatases are required to remove the phosphate, and the beginning of a new translational program results in the conversion of sporozoites into schizonts that are then able to infect the liver (Figure 2). In a very similar way, phosphorylation of Toxoplasma gondii’s eIF2 takes place in the transition between tachyzoite forms that convert into latent bradyzoites (Figure3).

The evolutionary distance between the eIF2 versions of apicomplexan parasites and humans, makes it possible to find differences in their regulation that could permit inhibition of the parasite translational machinery without producing the same effect in the human cells of an infected host. Although some kinases responsible for eIF2 phosphorylation in these organisms have been described, as well as the stresses that trigger the phosphorylation and activation of the different translational programs, it is still unknown which phosphatases produce the opposite effect. In any case, knowing the molecular processes that function as switches that activate the infective form of these parasites provides reliable targets for pharmacololgical inhibition at different levels.
References
- Zhang M, Joyce BR, Sullivan WJ, Jr., Nussenzweig V (2012) Translational Control in Plasmodium and Toxoplasma Parasites. Eukaryotic Cell. doi:10.1128/EC.00296-12 ↩
1 comment
[…] are quite easily directed by the control of only a small number of genes (those molecular switches we talked about here), it is crucial to define what those genes are activating or deactivating, in order to pursue the […]