How to describe hervibore teeth from their photographs
How to describe hervibore teeth from their photographs
Teeth are common elements in museum collections because they are better preserved than other parts of the body, being available for most living and fossil species. And they are extremely useful. Dental morphology has an eminent role in taxonomy, as systematics studies strongly rely on dental diagnostic characteristics.

Also, the enamel pattern of herbivore teeth and tooth crown height are regarded as phenotypic features with sound ecological effects and provide important information on the ability of individual species to cope with habitat conditions. Occlusal features of molars capture several aspects of the mechanics and durability of dental pieces during mastication and, being proxies for the diet and the environment of extinct species, have attracted a lot of interest in evolutionary and paleoecological studies.
One of the dental features that have received more attention is occlusal ‘dental complexity’. Dental complexity, complexity from here on, has been defined as any parameter measuring the number of features, ‘tools’ or ‘breakage sites’ on a tooth. The degree of enamel complexity is correlated with both body size and diet in different ungulate lineages: highly convoluted enamel on cheek teeth is linked with diets focused on higher abrasiveness and/or tough items like monocotyledonous grasses and/or grit.
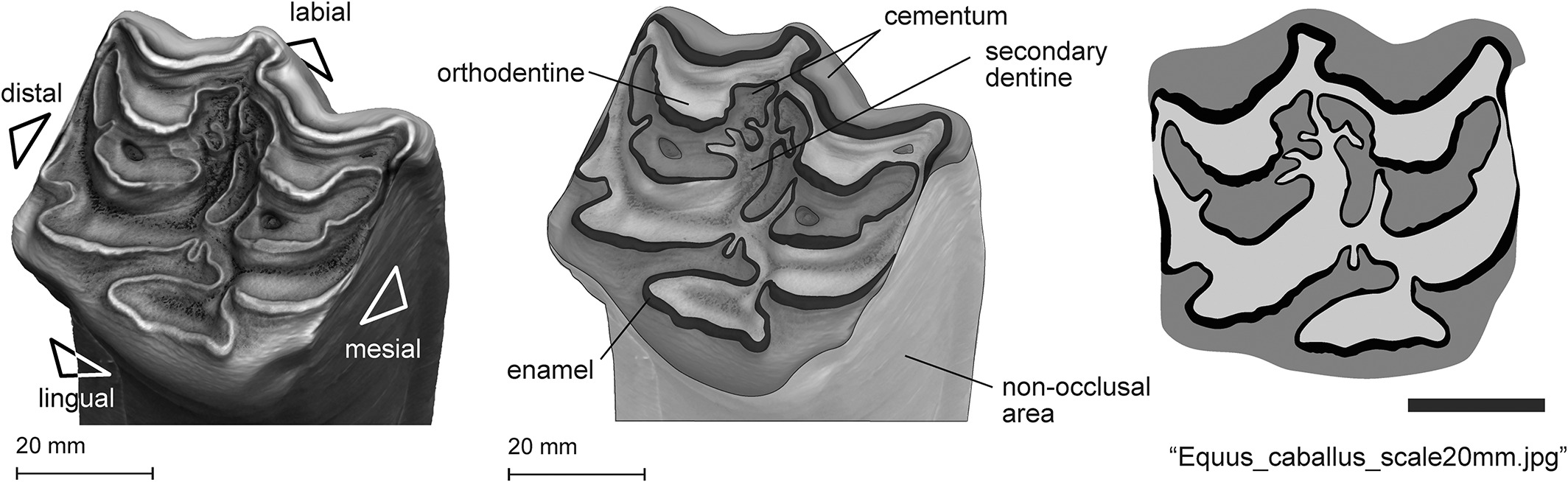
Folded is a new toolkit that provides 1 the instruments necessary to perform a comprehensive numerical description of the 2D occlusal patterns of herbivore mammal enamel. In other words, Folded has the potential to reconstruct the dietary variability, the mastication behaviour or the evolutionary adaptation in fossil and modern species using teeth photographs as an input.
A series of ‘topometric’ descriptors have been proposed for parametrizing tooth complexity from two-dimensional (2D) images. Folded uses three parameters: 2D orientation patch count, enamel folding, and local enamel thickness. In the case of 2D orientation patch count, adjacent linear segments with similar orientations are grouped into patches. Frequent changes in enamel orientation reflect complex tooth’s configurations.
Alternatively, orientation is quantified by means of enamel folding, defined as the inverse of ‘local coherency’. Coherency analysis has been demonstrated to be a reliable method of quantifying organic tissues, but has never been applied to dental enamel data. A folding index of zero would represent a completely straight, filamentary structure (such as a single, unbend enamel band) and tends to be one for pixels with no dominant orientation like in the case of sharp curves.
The method’s pipeline is automatic, requires traced 2D images as input and can process large numbers of specimens in seconds. The input can be obtained from images of worn molars in occlusal view (including the immense number of photographs/diagrams published in the literature), thus providing the opportunity for analysing vast amounts of data at virtually zero cost.
This approach will be of particular interest to the many institutions with extensive neontological or paleontological collections lacking access to 3D scanning devices or cutting-edge processing software, as well as research teams with limited budgets to visit museum collections.
Author: César Tomé López is a science writer and the editor of Mapping Ignorance
Disclaimer: Parts of this article may have been copied verbatim or almost verbatim from the referenced research paper/s.
References
- Sanisidro, O., Arganda-Carreras, I., & Cantalapiedra, J. L. (2023). Folded: A toolkit to describe mammalian herbivore dentition from 2D images. Methods in Ecology and Evolution, 14, 556– 568. doi: 10.1111/2041-210X.14042 ↩