Genetic architecture of autism is consistent across diverse populations
Genetic architecture of autism is consistent across diverse populations
A new study 1, co-led by researchers at the Icahn School of Medicine at Mount Sinai, demonstrates that genes associated with autism risk are largely the same across people of different ancestries. The findings, based on one of the largest genomic studies of Latin American individuals to date, provide strong evidence that the genetic architecture of autism is consistent across diverse populations. They underscore the importance of expanding genetic research beyond individuals of European ancestry.
Closing the ancestry data gap
Over the past decade, scientists have identified numerous rare genetic variants that confer substantial risk for autism and other neurodevelopmental disorders. However, most of these discoveries were made in cohorts composed predominantly of individuals of European ancestry, leaving open the question of whether autism’s genetic underpinnings differ across populations. This knowledge gap has contributed to disparities in genetic testing, including higher rates of inconclusive results among non-European individuals due to limited reference data.
To address this issue, the research team analyzed exome and genome sequencing data from more than 15,000 Latin American individuals across North, Central, and South America, including approximately 4,700 individuals diagnosed with autism. Latin American populations represent the largest recent mixed-ancestry group globally, with heritage that frequently includes Indigenous American, West African, and European origins. This rich genetic diversity provides a powerful opportunity to refine gene-disease associations, which can improve health outcomes for all populations.
What the researchers looked for
The study examined more than 18,000 genes for enrichment of rare, deleterious coding variants—genetic changes that can have immediate and profound clinical implications for diagnosis, treatment, and family counseling.
Consistent with prior research, rare, deleterious variants in highly conserved genes—genes that remain similar across species and populations over long periods of time—were disproportionately observed in individuals with autism. Researchers identified 35 genes significantly associated with autism in the Latin American cohort. These genes showed extensive overlap with those previously identified in genome-wide studies of individuals of European ancestry. The findings also provide support for several recently identified “emerging” autism-associated genes.
“Our results indicate that the core genetic architecture of autism is shared across ancestries,” said study senior author Joseph D. Buxbaum, Ph.D., Director of the Seaver Autism Center for Research and Treatment at Mount Sinai. “This suggests that the biology underlying autism is universal and reinforces the importance of ensuring that diverse populations are represented in genetic research.”
Revisiting gene conservation metrics
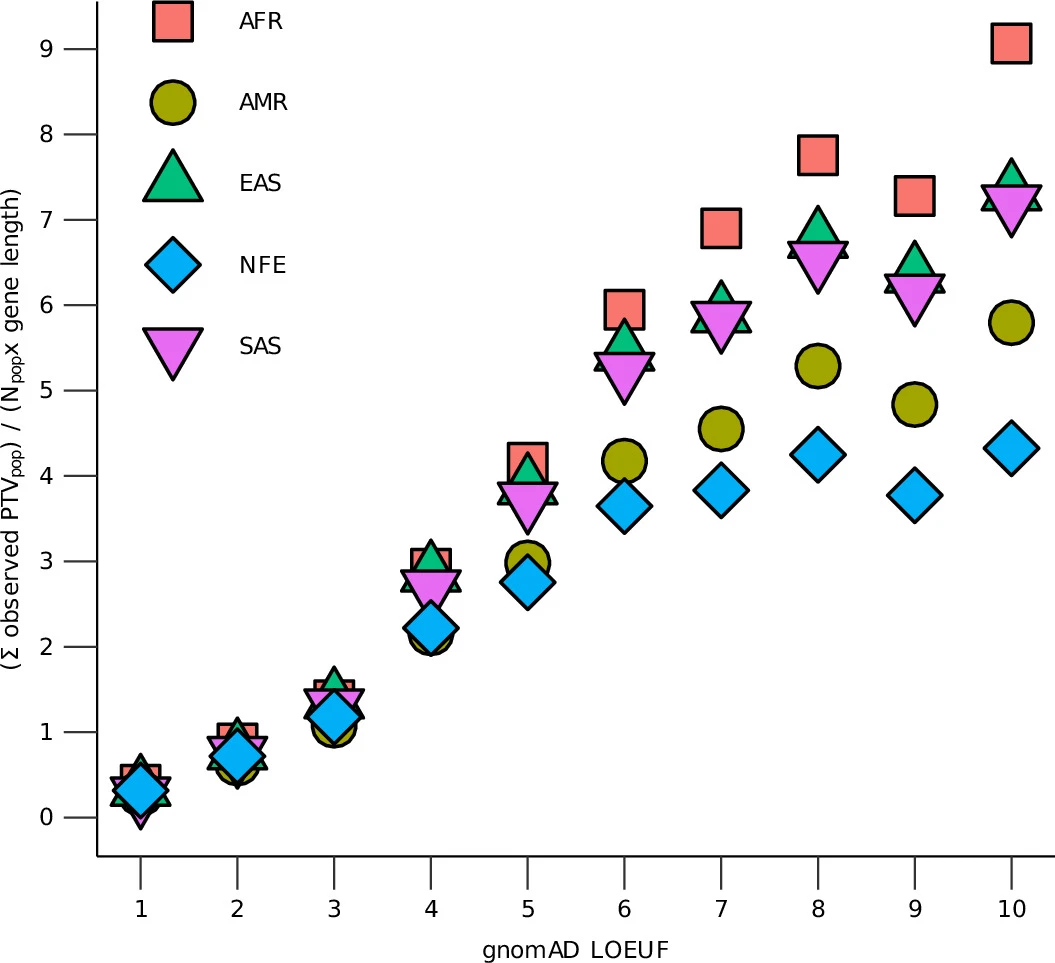
The study also evaluated widely used metrics that assess evolutionary conservation of genes, an important tool for prioritizing genes in clinical genetic analyses of neurodevelopmental disorders. The researchers found that these metrics, which were again largely derived from European-ancestry datasets, may overestimate conservation overall due to limited ancestral diversity in European populations. However, the metrics remain highly accurate for the most strongly conserved genes—including those most relevant to autism and other neurodevelopmental disorders.
The authors note that continued sequencing of diverse populations will further improve conservation metrics, particularly for less conserved genes, ultimately enhancing the accuracy of clinical genetic testing.
“These findings provide a road map for improving genetic diagnosis across ancestral groups,” said Dr. Buxbaum. “Expanding genomic research in underrepresented populations is essential to reducing health disparities and advancing precision medicine for autism and related conditions across all ancestral populations.”
The study’s results align with growing evidence that both rare and common genetic risk factors for complex disorders are shared across diverse populations. By demonstrating broad overlap in autism risk genes across ancestries, the research supports more inclusive approaches to genomic medicine and reinforces the universal biological foundations of autism.
References
- Natividad Avila, M., Jung, S., Satterstrom, F.K. et al. (2026) Deleterious coding variation associated with autism is shared across ancestries. Nat Med doi: 10.1038/s41591-026-04228-6 ↩