Can drugs become toxic when exposed to light?
Can drugs become toxic when exposed to light?
Author: Neus Aguilera-Porta is a Ph.D. student (ITN-EJD-TCCM) at Universidad Autónoma de Madrid
Medicines are a mixture of pharmaceutically active molecules with excipients, designed with the aim of healing, relieving the pain with as minimal side effects as possible. Having this in mind, could the properties of a molecule be modified when absorbing a light photon? Could then these drugs lose their potency as a pharmaceutical compound?
The photostability of a therapeutic compound is a must in order to ensure that its properties will not be modified over its lifetime, from formulation and manufacture through its active life cycle, including handling, packaging, and dispensation. As the absorption of an UV-visible light photon is energetically comparable to the bond energy in an organic compound its absorption could induce undesirable changes in the active principle structure. Thus, it is crucial to investigate the photophysics and the photoreactive paths activated upon photon absorption to monitor the potency of pharmaceutical compounds and to identify potential side products of photochemical origin 1.
Firsts attempts to predict photostability and phototoxicity of drugs and pharmaceutical excipients were based on the calculation of the frontier orbitals HOMO-LUMO energy gap are available. However, their predictive power is limited and therefore they are sometimes complemented with indicators of light absorption intensity, i.e. the molar extinction coefficient, ε.2
Molecular modelling offers a broad spectrum of tools that can potentially result very useful to the pharmaceutical industry in the prediction of the photophysical and photochemical intrinsic behaviour of drugs upon light absorption and thus to serve as a first step towards a better assessment of its quality and safety.
This Jablonski diagram in Figure 1 schematically summarizes all the possible photophysical processes which are likely to occur in a molecule after the absorption of an UV-visible light photon leading eventually to its deactivation. The absorption of a light photon may bring the molecule from the electronic ground to an excited state in the Franck-Condon (FC) region. The lowest lying electronic excited states are the result of promoting a valence electron from an occupied to an unoccupied orbital.
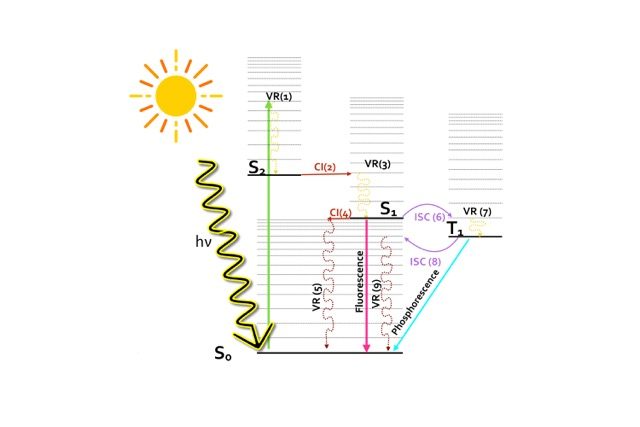
Let’s say that by absorbing a light photon of the appropriate energy, the molecule is excited from the ground state configuration (S0) to the second excited state (S2). This energetic state, S2, could decay back to the ground state via different competing photophysical processes.
For instance, the system could undergo an initial radiationless deactivation via a vibrational relaxation (VR1) to decay to the lowest vibrational state of S2 and an internal conversion (IC) through conical intersection (CI 2) to populate the first excited state. The S1 can then further decay back to the ground state following different radiative or non-radiative photophysical processes respectively, such as fluorescence (radiative) or IC through conical intersection (CI 4).
However, it might occur that the singlet and the triplet surfaces cross each other at an intersystem crossing point (ISC), depicted as ISC 6 in Figure 1, allowing the transfer of population to the triplet state. From this lower triplet, the excited system could either decay to the ground state radiatively, via phosphorescence, or otherwise radiationless, via the ISC 8 funnel.
Besides the above mentioned photophysical processes, there are other photochemical processes which might also occur and which are not represented in the Jablonski diagram. If the excited states would live for enough time, they could evolve to other species or alternatively could react with other molecules from the environment leading to the formation of photoproducts.
Leaving aside, the photochemical reactions that could compete with the photophysical processes, exploring the potential energy surface (PES) of drugs with computational approaches could result very useful to map the possible relaxation pathways of photo-excited drugs. However, PES are extremely complex mathematical objects, with a high dimensionality, 3N-6 or 3N-5, where N is the number of atoms in the molecule. This complexity transfers us from the previous Jablonski one-dimensional “staircase” scheme, to an entangled landscape comparable to Escher’s “Relativity” lithography.
To simplify this task a common practice is to perform minimum energy path calculations starting from the FC region, transition states, or singlet-singlet and singlet-triplet crossing points to constrain the study of regions of the PES which are energetically relevant. These calculations often result in 2D or 3D simplified profiles, where the potential energy is represented against one global coordinate or few relevant degrees of freedom to the deactivation mechanism.
Among all possible competing deactivation processes, the most probable(s) can be determined undertaking molecular dynamics simulations that statistically estimates the probabilities of the different deactivation mechanisms.
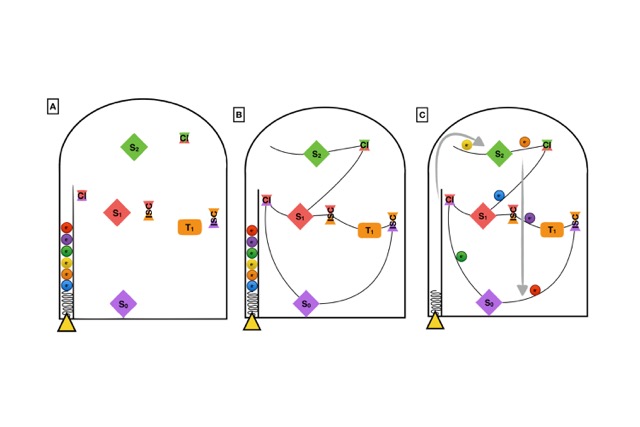
In order to better illustrate this combined static and dynamic protocol, in the following we will establish an analogy with the pin ball board game. Let’s consider the geometrical targets, in Figure 2, to be the stationary and crossing points in a PES (A), localized with static quantum chemical methods. The MEPs would be defined by the imaginary lines connecting these targets (B). The most probable deactivation pathways would be determined after launching a set of balls in the game and statistically stablishing which are the preferred paths followed to decay back to the ground state (C).
The insight provided by molecular modelling into the photophysical and photochemical properties of compounds can be profited to assist in the design of drugs with controlled side effects. Unfortunately, these calculations cannot be directly transferred to industry due to their high cost. So there is urgent need to find other indicators that could help anticipating the photophysical properties of drugs, and thus to develop current phototoxicity models to predict the inherent photosensitive properties of active principles.
Aknowledgements
This project was supported by the European Union’s Horizon 2020 research and innovation programme under the Marie Curie Skłodowska-Curie grant agreement (Nº 642294).
The computations were performed with the resources provided by Centro de Computación Científica at Universidad Autónoma de Madrid (CCC-UAM).