Stromal senescence drives therapy resistance in T-ALL
Stromal senescence drives therapy resistance in T-ALL
Author: Marta Irigoyen is a postdoctoral researcher at CIC bioGUNE
T-cell lymphoblastic lymphoma (T-LBL) is a precursor T cell lymphoma. T-LBL represents the second most common subtype of Non-Hodgkin lymphoma in both children and adolescents; however, it is less common in adults 1. T-LBL share many characteristics with the precursor T cell leukemia (T-ALL), however, various studies have indicated that T-ALL and T-LBL might be genetically distinct due to different gene abnormalities 2. Indeed, despite the similarities between these two entities, T-LBL often presents clinically with a large mediastinal mass and rarely involves the bone marrow, unlike T-ALL, which often involves the bone marrow. Regarding the survival rate, both T-ALL and T-LBL have a high 5-year survival rate in children but, in adults, the overall 5-year survival rate is less favorable. Besides, despite a comprehensive treatment regime, a high percentage of patients, especially adults, relapse and acquire therapy resistance 3. Up to day, few studies have addressed the potential mechanisms leading to therapeutic resistance in T-LBL/ALL. The majority of these studies have indicated an important role of the microenvironment in providing pro-survival signals to the leukemic cells 4. However, the role of stromal cells in the survival and therapeutic resistance of the leukemic cells has not been explored yet. For this reason, in the present paper Habiel and colleges aimed to study the relevance of these stromal cells in T-ALL 5.
Firstly, in order to analyze in more depth the differences between these two leukemic cells in their respective microenvironments, authors examined publicly available gene expression arrays comparing lymphatic infiltrated T-LBL to bone marrow infiltrated T-ALL cells 6 and performed canonical pathway analysis. Thus, data showed that there was a marked enrichment of pro-fibrotic transcripts in T-LBL relative to T-ALL and TGFß signaling pathway was the top most predicted activated upstream regulator in T-LBL relative to T-ALL. Since they observed evidences of fibro-proliferation and remodeling in T-LBL lymphatic biopsies, to continuous they work they selected CEM cells because when were intravenously administered into mice induced a disease with similar characteristics to an adult T-LBL and, only a few infiltrating of CEM cells were detected in the bone marrow of mice 7. To determine whether CEM cells induced fibro-proliferation in mice, they performed Masson’s trichrome staining to stain and visualize the collagen deposits in various CEM-infiltrated organs. Interestingly, Masson’s trichrome staining was increased in lungs, heart and kidneys but not in the spleen or the liver respect to control mice. Next, having into account that T-LBL frequently is present together with pleural effusions, they induced a pulmonary injury in the lung of mice that received CEM cells through bleomycin (a chemoterapeutic agent) administration via oropharyngeal aspiration. Interestingly, hydroxyproline levels (hydroxyproline is one of the most relevant markers in collagen metabolism) were significantly increased in the lungs of mice challenged with both the CEM + Bleo cohort compared with bleomycin alone, CEM alone, or control mice. According with this results, the increased collagen staining was also evident in the CEM + Bleo group compared with the other groups. Besides, CEM infiltrates were detected in the lung, liver, and spleen but not in the kidney or the heart after CEM + bleo or CEM challenge.
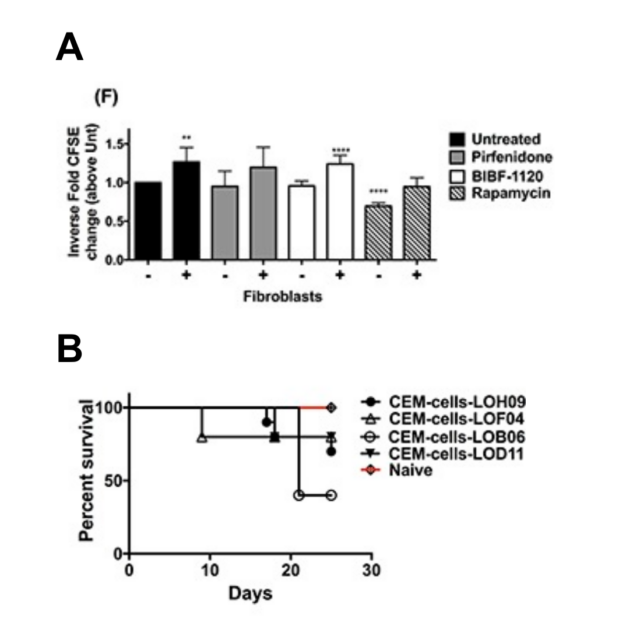
Given these findings, they next examined the interplay between fibroblasts and leukemic cells using various in vitro techniques. Thus, they generated primary lung stromal cells or fibroblasts from biopsies or explants and co-cultured CEM cells with them. After 21 days of co-culture, they observed a phenotypic divergence of CEM cells co-cultured with fibroblasts increasing their cell size and morphology. To determine whether these changes were able to alter their therapeutic sensitivity and efficacy, the authors performed various studies in which CEM cells were co-cultured in the presence or absence of fibroblasts and various therapeutics. Interestingly, fibroblasts rendered CEM cells more resistant to the different chemoterapeutic agents. To assess whether stromal cells could propagate persistent changes in CEM cells, the authors co-cultured CEM cells with stromal cells and then transferred them into fresh culture dishes. After a minimum of 48 hours, the cells were treated with various therapeutics. The stromal cell–induced changes in CEM cells were maintained, resulting in different levels of resistance to multiple chemotherapeutic agents compared with CEM cells that had not been co-cultured with human stromal cells.
To further validate the phenotypic divergence of co-cultured CEM cells and effects of stromal cells on CEM induced lymphoma/leukemia in mice, CEM cells (after co-culture with stromal cells) were intravenously administered into mice. At 25 to 27 days post-injection, injected human T-cells were histologically detected in spleen, lung, heart, liver and bone marrow in mice. However, mice injected with CEM cells co-cultured with the LOF04 fibroblast line showed much less invasion into spleen, lung, liver and bone marrow and a better survival as compared with other CEM cells conditioned with additional fibroblast cell lines and similarly introduced into mice. To identify the differential effects of stromal cells on CEM cells, further analysis of LOF04 fibroblasts were undertaken. Interestingly, this stromal cell line was non-proliferative and senescent because presented a flattened morphology, expressed higher ß-galactosidase activity, produced more reactive oxygen species (ROS) and showed significantly higher transcript levels for cell cycle checkpoint proteins relative to proliferating fibroblasts. Given the marked genotypic divergence observed when CEM cells were cultured with senescent versus proliferating fibroblasts, authors aimed to further characterize potential molecular mechanisms leading to this difference.
It is well known the role of BRCA1 in anti-oxidant responses and the Mismatch Repair (MMR) Pathway in the maintenance of microsatellites. Taking these data into account, authors performed various experiments to analyze the expression of components from these pathways in CEM cells after co-culture with fibroblasts and they found a marked reduction in the expression of components of the MMR and BRCA1 in CEM cells co-cultured with fibroblasts compared with CEM cells prior to co-culture. Since micro-RNAs (miRNAs) have been shown to modulate DNA repair pathways, they hypothesized that miRNA-directed mechanisms via secreted microvesicles might lead to the specific modulation of the MMR and BRCA1 pathways in the co-cultured CEM cells. According with this hypothesis, the expression of the exosomal marker CD63 was higher on stromal cell-conditioned CEM cells compared with control CEM cells. To determine whether MMR and BRCA1 targeting miRNAs were present in senescent stromal cell derived microvesicles, they analyzed the RNA from microvesicles looking for DNA repair targeting miRNAs. Remarkably, mature MMR and BRCA1 targeting miRNAs were abundantly expressed in fibroblast-derived microvesicles and were increased in CEM cells co-cultured with senescent stromal cells.
Collectively, these results suggest that in addition to generating oxidative radicals, senescent fibroblasts release exosomes containing miRNAs targeting the MMR and BRCA1 pathways potentially contributing to the divergence of leukemic cells located in their microenvironment.
References
- Burkhardt B, Zimmermann M, Oschlies I, Niggli F, Mann G, Parwaresch R, Riehm H, Schrappe M et al. (2005) The impact of age and gender on biology, clinical features and treatment outcome of non-Hodgkin lymphoma in childhood and adolescence. Br J Haematol. 2005;131:39-49. PMID: 16173961 DOI: 10.1111/j.1365-2141.2005.05735.x. ↩
- Raetz EA, Perkins SL, Bhojwani D, Smock K, Philip M, Carroll WL, Min DJ. Gene (2006) expression profiling reveals intrinsic differences between T-cell acute lymphoblastic leukemia and T-cell lymphoblastic lymphoma. Pediatr Blood Cancer. 2006;47:130-40. PMID: 16358311 DOI: 10.1002/pbc.20550. ↩
- Bonn BR, Rohde M, Zimmermann M, Krieger D, Oschlies I, Niggli F, Wrobel G, Attarbaschi A et al. (2012) Incidence and prognostic relevance of genetic variations in T-cell lymphoblastic lymphoma in childhood and adolescence. Blood. 2013;121:3153-60. PMID: 23396305 DOI: 10.1182/blood-2012-12-474148. ↩
- Sun Y. Tumor (2015) microenvironment and cancer therapy resistance. Cancer Lett. 2015 PMID: 26272180 DOI: 10.1016/j.canlet.2015.07.044. ↩
- Habiel DM, Krepostman N, Lilly M, Cavassani K, Coelho AL, Shibata T, Elenitoba-Johnson K, Hogaboam CM. Senescent stromal cell-induced divergence and therapeutic resistance in T cell acute lymphoblastic leukemia/lymphoma. (2016) Oncotarget. 2016;7:83514-83529. PMID: 27835864 DOI: 10.18632/oncotarget.13158. ↩
- Mussolin L, Holmes AB, Romualdi C, Sales G, D’Amore ES, Ghisi M, Pillon M, Rosolen A et al. (2014) An aberrant microRNA signature in childhood T-cell lymphoblastic lymphoma affecting CDKN1B expression, NOTCH1 and growth factor signaling pathways. Leukemia. 2014;28:190-12. PMID: 24727675 DOI: 10.1038/leu.2014.134. ↩
- Hosler GA, Bash R, Scheuermann RH. (2000) Kinetics of early therapeutic response as measured by quantitative PCR predicts survival in a murine xenograft model of human T cell acute lymphoblastic leukemia. Leukemia. 2000;14:1215-24. PMID: 10914545 DOI: 10.1038/sj.leu.2401814. ↩