Unraveling protein knots

Proteins are sophisticated machines that perform most of our cellular activities. Proteins are codified in the cellular DNA and synthesized as linear chains that need to get properly folded in a 3D structure to become functional. This state is named native structure.
Several perturbations can challenge the native structure of the proteins converting them into dysfunctional and misfolded proteins. In this misfolded state, proteins present “sticky” patches that bind unspecifically to other proteins, forming a huge net of protein aggregates, a process that is highly toxic to cells.
Among the perturbations that lead to protein aggregation are the oxidative stress, heat shock, mutations and aberrant protein turnover.
In humans several diseases are associated with increased protein aggregation, notably Alzheimer. However, little is known about how human or other animal cells deal with those protein aggregates. In lower organisms like bacteria, plants or fungi, a network form by Hsp70 and Clp proteins are able to unravel the protein aggregates turning them back into functional proteins. However animal cells lack Clp proteins, essential partners of this network, raising several questions: Are the animal cells able to unravel protein aggregates? Which proteins perform that activity?
In 2006 Cohen and colleagues demonstrated1 that animal cells, under certain stimuli, are able to unravel/disaggregate the protein aggregates form by Ab42, a notorious precursor of Alzheimer disease. However the proteins responsible for this activity were unknown.
A few months ago, the group of Prof Bernd Bukau from Heidelberg (Germany) published a key study2 in EMBO Journal that unveiled the identity of the protein network that repairs protein aggregates in animals.
The authors assumed that the repair network in animal cells would use Hsp70 proteins, a highly conserved group of proteins that assist protein folding in all known organisms3. A surprising finding of this study was that animal cells do not require a protein similar to the bacterial or fungi Clp to perform the activity. Instead Hsp70 proteins perform the disaggregation process in coordination with a poorly study group of related proteins named Hsp110 proteins (Apg2 in humans).
In their paper, the authors demonstrate that Hsp70 and Hsp110 proteins are able to recover functional proteins from aggregates in vitro, in an animal model and in human cultured cells.
In lower organisms Hsp70 proteins initiate the aggregate reactivation. They are followed by Clp proteins, which bring the partially reactivated proteins through an internal hole, like a thread through a needle, in a process that is not fully understood (Winkler et al, 2012). According to the author’s data, in animals Hsp110 proteins assist to the Hsp70 enzymatic activity, which is the sole carrier of the reactivation process. The evidence presented by the authors suggests that Hsp110 proteins regulate the energy flux of Hsp70 proteins (nucleotide exchange). However the molecular mechanism that enables Hsp110 proteins to transform Hsp70 proteins into disaggregating machines is still unclear.
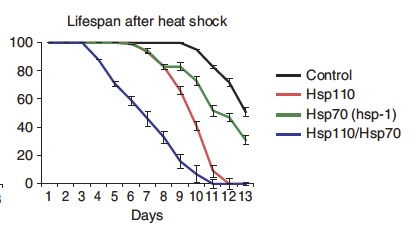
The authors also provided evidence of Hsp110 proteins localize together with protein aggregates in human cells. Moreover they showed that animal models (C. elegans) lacking Hsp110 activity were unable to reactivate protein aggregates and had reduced lifespan upon heat shock (Figure 2), a well known cellular stress that leads to protein aggregation.
Overall, the scientists described a novel protein network in animal cells that is able to reactivate protein aggregates. Whether these proteins are involved in the disaggregation of disease-associated protein aggregates in humans is still unclear. To this point is important to note that a member of the Hsp110 protein family has been described4 to localize together with a specific type of protein aggregates named amyloid inclusions, which, as described above, play a critical role in Alzheimer’s disease progression.
Thus, the author’s discoveries offer a new approach to study protein aggregate-related diseases. Further studies may shed light on the cellular mechanisms that prevent and unravel those undesirable protein knots in the context of human disease.
References
- Cohen E, Bieschke J, Perciavalle RM, Kelly JW, Dillin A (2006) Opposing activities protect against age-onset proteotoxicity. Science313(5793):1604-10. ↩
- Rampelt H, Kirstein-Miles J, Nillegoda NB, Chi K, Scholz SR, Morimoto RI, Bukau B (2012) Metazoan Hsp70 machines use Hsp110 to power protein disaggregation. EMBO J31(21):4221-35. ↩
- Winkler J, Tyedmers J, Bukau B, Mogk A (2012) Chaperone networks in protein disaggregation and prion propagation. J Struct Biol 179(2):152-60. ↩
- Olzscha H, Schermann SM, Woerner AC, Pinkert S, Hecht MH, Tartaglia GG, Vendruscolo M, Hayer-Hartl M, Hartl FU, Vabulas RM (2011) Amyloid-like aggregates sequester numerous metastable proteins with essential cellular functions. Cell144(1):67-78. ↩
8 comments
Nice article. I am surprised that misfolded proteins are beginning to become one of the most important contributors to many pathologies associated to cell death, like oxidative stress a few years ago. It looks like both oxidative stress and accumulation of misfolded proteins are cooperative processes that enhance a feedback affecting many other cellular processes and probably is what finally unbalances the cell conditions towards a pathologic state, independently of what initial damage triggers the accumulation. Interesting…
Very interesting. Although I do not question that some protein aggregates may be toxic, some controversy exists about whether it can be a protective mechanisms against other misfolded proteins, such as the proteins with toxic gain-of-function mutations (e.g. huntingtin).
I think it would be interesting to see what could happen in animal or cell models of Huntington’s disease if Hsp110 is silenced. Would it worsen the phenotype?
What it is also interesting is that dividing cells seem to be able to “deal” with protein aggregates by leaving them in just one of the daughter cells after mitosis, creating a sort of “doomed” cell that would probably die while the other daughter cell will remain clean. So, what happen in cells that do not divide?? Well, just check what happens in mature neurons: they accumulate unprocessed + misfolded + non-degraded proteins that contribute to neuronal diseases.
Sure, that is a very good point; and again it is somehow similar to what happens with ROS accumulation: dividing cells more easily deal with ROS detoxification and its products, whereas quiescent or non-dividing cells at some points are overwhelmed by the accumulation of ROS and deleterious oxidation products and finally die.
Life is really difficult for cells, if they don’t divide they accumulate garbage and bad things, however if they divide too quick and lose control, they become cancer cells. Is accumulation of misfolded proteins a triggering event of apoptosis in order to avoid cancer, in a quality control kind of selectivity? The chicken and the hen, again… ;D
There is evidence on mutations, which drive certain proteins to unfunctional and unfolded/misfolded states, and are linked to tumorigenesis ( http://www.sciencedirect.com/science/article/pii/S0968000400891008 ; I think some p53 mutations work like that). No idea whether increased aggregation can cause defects on the DNA copying or repairing machinery and thus, rise new mutations that could lead to cancer. It is really intriguing!!!!!!
@moigaren I agree, indeed if you see how I wrote it ” In this misfolded state, proteins present “sticky” patches that bind unspecifically to other proteins, forming a huge net of protein aggregates, a process that is highly toxic to cells.”
The process is toxic because the sequestering/damaging of cellular components but there is growing evidence that the product, i.e. the aggregate, is more like a final, semi-inert, compartment. Something like “The bullet that kills you is the one which is moving, but you can put as many as you want in a box”
At the time I started my PhD in 2004 aggregates were considered quite dangerous. With time the idea was refined and the aggregation process itself got the attention while the final deposits were considered more like a deposit by many scientists
[…] stress produced by the accumulation of unfolded proteins (a topic reviewed in Mapping Ignorance, here and here). Now the landscape became enriched with several new data, and with the work we review […]
Opioid analgesics have also been proven by clinical evidence to be very effective in alleviating the type of chronic pain commonly
experienced by those suffering from AS, especially in time-release formulations.
Also, you can order cucurmin, the ingredient in turmeric being studied, from most
health food stores or online for example, from
the Life Extension Foundation. The question is will it also help humans in the early
stages of Alzheimer’s disease.