The power of herbaria: a time machine for plant biology research
The power of herbaria: a time machine for plant biology research
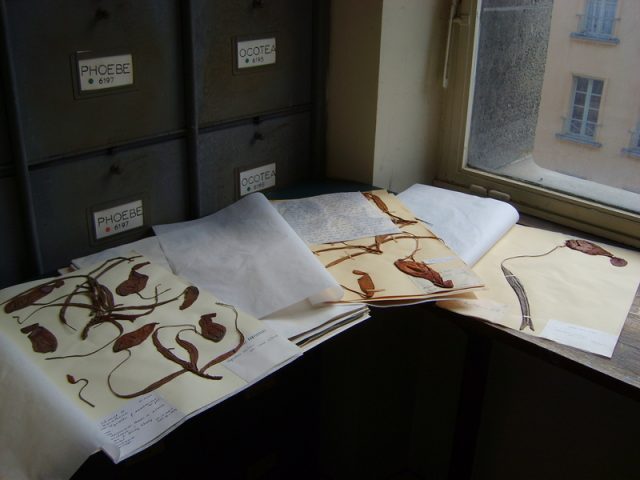

Naturalists and scientists have been collecting plants or plant parts during centuries to make collections and catalogues known as herbaria (sing. herbarium) that have been traditionally used for comparative taxonomy and systematics research. The first herbarium collections were compiled along with the foundation of botanical gardens during the first half of the 16th century. The oldest preserved herbarium is located in Rome and was begun by Gherardo Cibo in 1532. Today there exist around 3000 active herbaria with more than 350 million specimens stored worldwide (Figure 1).
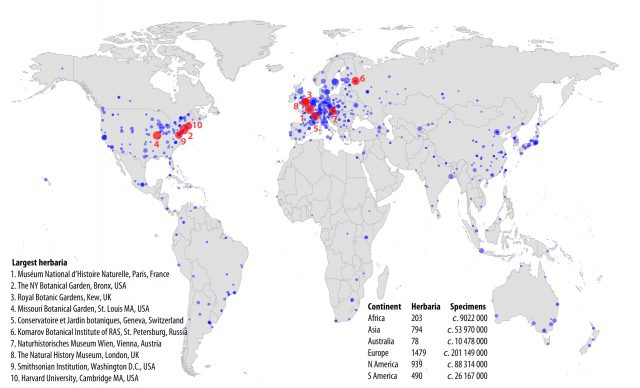
Each herbarium specimen represents an image of the conserved plant and is accompanied by metadata which include geographical location, date of collection and sometimes additional information such as parts that were lost during pressing, life conditions, smell, etc. These metadata are extremely valuable and essential to properly interpret the results obtained from herbaria. Today, herbarium specimens are being used for applications that go far beyond the classical taxonomy studies and may be employed purposes that could not have been imagined by their collectors 1. Figure 2 illustrates some of these applications such as studying historical species distribution, plant adaptation to climate change, ancient contamination, etc. These studies may be non-invasive or they may need to destroy part of the sample such us for obtaining DNA of for analyzing the chemical composition of the samples.
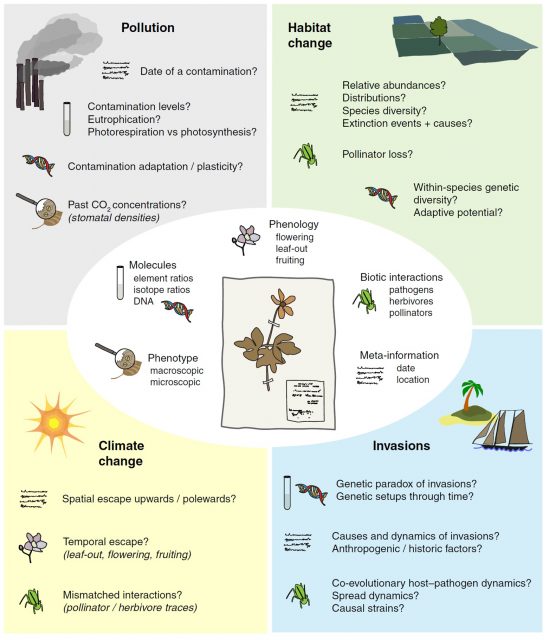
Here below I will summarize some examples of studies that have used herbarium specimens for different purposes to highlight the potential of this amazing resource:
Phenology is the study of the relationship between climate and life cycle, such as flowering in the case of plants. The study of phenology is vital to understand how plants will respond to climate change. Interestingly, herbarium specimens represent snapshots of phenological events and thus, the use of herbaria is a great opportunity. For example, Calinger et al. 2 studied 5053 herbarium specimens from 1985 to 2015 belonging to 141 species of midwestern United States and Davis et al. 3 analyzed 1108 herbarium records belonging to 20 species from Massachusetts herbaria collected between 1852-2012. Both works analyzed flowering occurrence and suggested earlier flowering with increasing temperature corresponding to 3.5 days/ºC (Davis et al., 2013) and 2.4 days/ºC (Calinger et al., 2013). Importantly both works underline the great opportunity of using herbaria to study global climate change.
Herbarium specimens may be also used to study contamination; for instance, by metal pollution. As examples, Peñuelas and Filella 4 studied specimens from 24 species collected in Spain during four different decades (1920-30; 40-50; 60-70 and 85-95) and Rudin et al. 5 used specimens from 1846 to 1916 from Rhode Island. In both works the accumulation of heavy metals in plant tissues was analyzed. Although the results showed variability in function of the species, one of the common observations of both studies was a reduction in lead concentration in most recent samples probably because of the decline in industrial activity and thanks to the environmental legal regulations. Besides, these studies highlight the potential of using herbarium specimens to study contamination.
During the last decade, the huge progress of sequencing technology has greatly facilitated the use of specimens from herbaria to study different aspects of plant evolution. For example, Exposito-Alonso et al. 6 used Arabidopsis thaliana species as a model to study plant colonization. The authors sequenced the genomes of 73 modern specimens but also of 23 specimens from herbaria of North America, the oldest one being from 1863 and with at least one sample per decade starting from the oldest record. The results obtained allowed the authors to establish the initial colonization event about 400 years ago. Moreover, they found a number of mutations conserved in the genome modern specimens that were linked to important traits of ecological importance, such as flowering time and therefore, could be key for recent plant adaptive evolution.
Finally, another example of using herbaria in combination with sequencing technology is the work of Yoshida et al. 7 who focused on studying Phytophtora infestans, the pathogen in charge of potato late blight disease, which became famous for having triggered the Irish Great Famine in 1840. Yoshida et al., sequenced the genome of 15 modern strains and also 11 strains collected from herbarium samples of infected potato and tomato leaves collected from continental Europe, Great Britain, Ireland, and North America in the period from 1845 to 1896, and were able to conclude that the 19th century epidemic was caused by a single P. infestans genotype which persisted for at least 50 years and that is from the examined modern strains.
I hope that from today when you think about a herbarium the image of old folders with dry plants full of dust disappears from your mind and is replaced by an amazing and powerful tool that can be used to understand many aspects of ancestral and modern plant biology.
References
- Lang PL, Willemns FM, Scheepens JF, Burbano HA, Bossdorf O (2018) Using herbaria to study global environmental change. New Phytologist doi.org/10.1111/nph.15401 ↩
- Calinger KM, Queenborough S, Curtis PS (2013) Herbarium specimens reveal the footprint of climate change on flowering trends across north-central North America. Ecology Letters 16: 1037-1044. ↩
- Davis CC, Willis CG, Connolly B, Kelly C, Ellison AM (2015) Herbarium records are reliable sources of phonological change driven by climate and provide novel insights into species’ phenological cueing mechanisms. American Journal of Botany 102: 1599–1609. ↩
- Peñuelas J and Filella I (2002) Metal pollution in Spanish terrestrial ecosystems during the twentieth century. Chemosphere46: 501–505. ↩
- Rudin SM, Murray DW, Whitfeld3 TJS (2017) Retrospective analysis of heavy metal contamination in Rhode Island based on old and new herbarium specimens. Applications in Plant Sciences 5(1): 1600108. ↩
- Exposito-Alonso M, Becker C, Schuenemann VJ, Reiter E, Setzer C, Slovak R, Brachi B, Hagmann J, Grimm DG, Chen J, Busch W, Bergelson J, Ness RW, Krause J, Burbano HA, Weigel D (2018) The rate and potential relevance of new mutations in a colonizing plant lineage. PLoS Genetics 14(2):e1007155. ↩
- Yoshida K, Schuenemann VJ, Cano LM, Pais M, Mishra B, Sharma R, Lanz C, Martin FN, Kamoun S, Krause J, Thines M, Weigel D, Burbano HA (2013) The rise and fall of the Phytophthora infestans lineage that triggered the Irish potato famine. Elife 2:e01108. ↩
4 comments
[…] Herbarioak, XVI. mendearen amaieran egiten hasi zieren landare bildumak, iraganera biadaiatzeko eta gaur egun garrantzitsua den informazioa biltzeko tresna bilakatu dira. Daniel Marinoren artikulua: The power of herbaria: a time machine for plant biology research. […]
[…] Los herbarios, esas colecciones de plantas que empezaron a confeccionarse en el siglo XVI, se han convertido en herramientas con las que viajar al pasado para recabar información relevante hoy día. Daniel Marino en The power of herbaria: a […]
Will it be available as a PDF on-line?
[…] The power of herbaria: a time machine for plant biology research (15/10/2018). Daniel Marino. […]